
The flowerpot snake (Indotyphlops braminus), known as the world’s only obligate parthenogenetic snake, reproduces solely via all-female parthenogenesis. Widely dispersed across tropical and subtropical regions largely due to inadvertent transport in potted plants, this diminutive snake has long puzzled biologists with its unusual reproductive mode. Now, in a study published in Science Advances, researchers led by Professor Jia-Tang Li, from the Chengdu Institute of Biology, Chinese Academy of Sciences, have assembled a high-quality, chromosome-level genome and partitioned it into three distinct subgenomes (A, B, and C). revealing its unique triploid parthenogenetic reproductive mechanism and shedding new light on the evolution of polyploidy in vertebrates.
Using multi-omics techniques—including long-read PacBio sequencing and Hi-C chromosomal mapping—the team revealed that the ancestral genome underwent a rapid speciation event approximately 41 million years ago. This event, accompanied by chromosomal fusion, resulted in a unique triploid genomic architecture that underlies the snake’s parthenogenetic reproduction.
The flowerpot snake’s genome comprises 42 chromosomes that can be divided into three subgenomes. Detailed analysis uncovered that two of the chromosomes appear in duplicate relative to previous karyotypic reports—a finding that resolved longstanding discrepancies in chromosome counts.
Comparative genomic analyses with related diploid snake species identified a fusion event in the ancestral genome, leading to distinct subgenome partitioning. This rearrangement may play a crucial role in facilitating compatibility among hybrid genomes.
The researchers observed extensive pseudogenization in genes associated with immune response and spermatogenesis. Such genetic alterations may reduce inter-subgenome immune conflicts, thus ensuring the viability of parthenogenetic reproduction.
Transcriptomic and epigenetic analyses of ovarian tissues revealed markedly elevated expression of key DNA repair genes—such as ALKBH2 and MDC1—suggesting that premeiotic endoreplication may help overcome challenges during meiosis in triploid organisms.
By uncovering the genomic mechanisms behind the flowerpot snake’s parthenogenesis, this study challenges traditional views that asexual reproduction inevitably leads to an evolutionary dead end. Instead, it provides a new perspective on how allopolyploid species may maintain genetic diversity and adapt to diverse environments without the need for male contribution.
The detailed dissection of the flowerpot snake genome not only reveals the evolutionary innovations that support parthenogenesis but also offers a paradigm for understanding polyploid evolution in vertebrates. Our findings pave the way for further investigations into the genomic and reproductive dynamics of asexual species.
Figure Captions

Figure 1. Flowerpot snake in its natural habitat. (Photo courtesy of Miu Jingling)
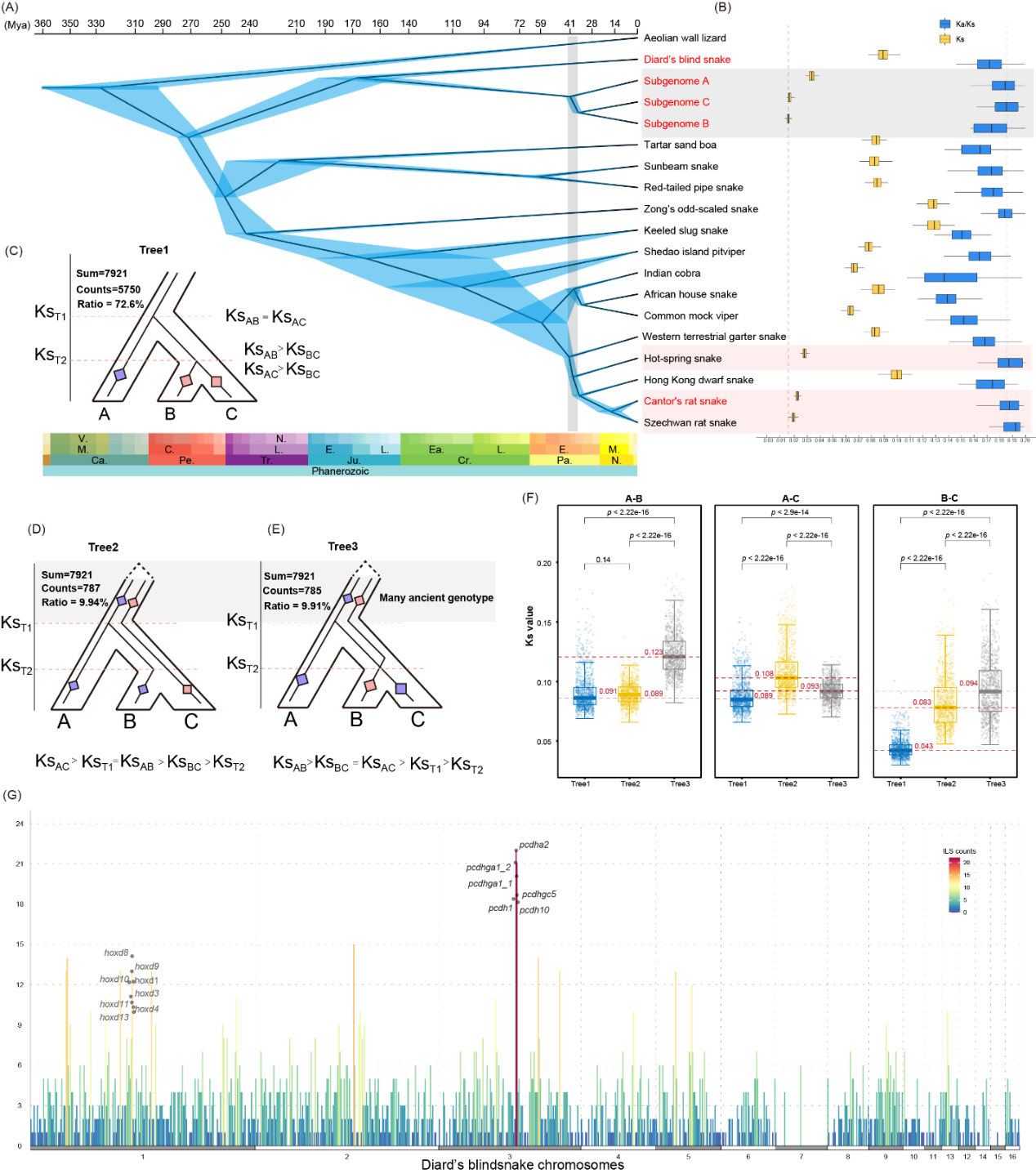
Figure 2. Schematic of the evolutionary history of the flowerpot snake's subgenomes.
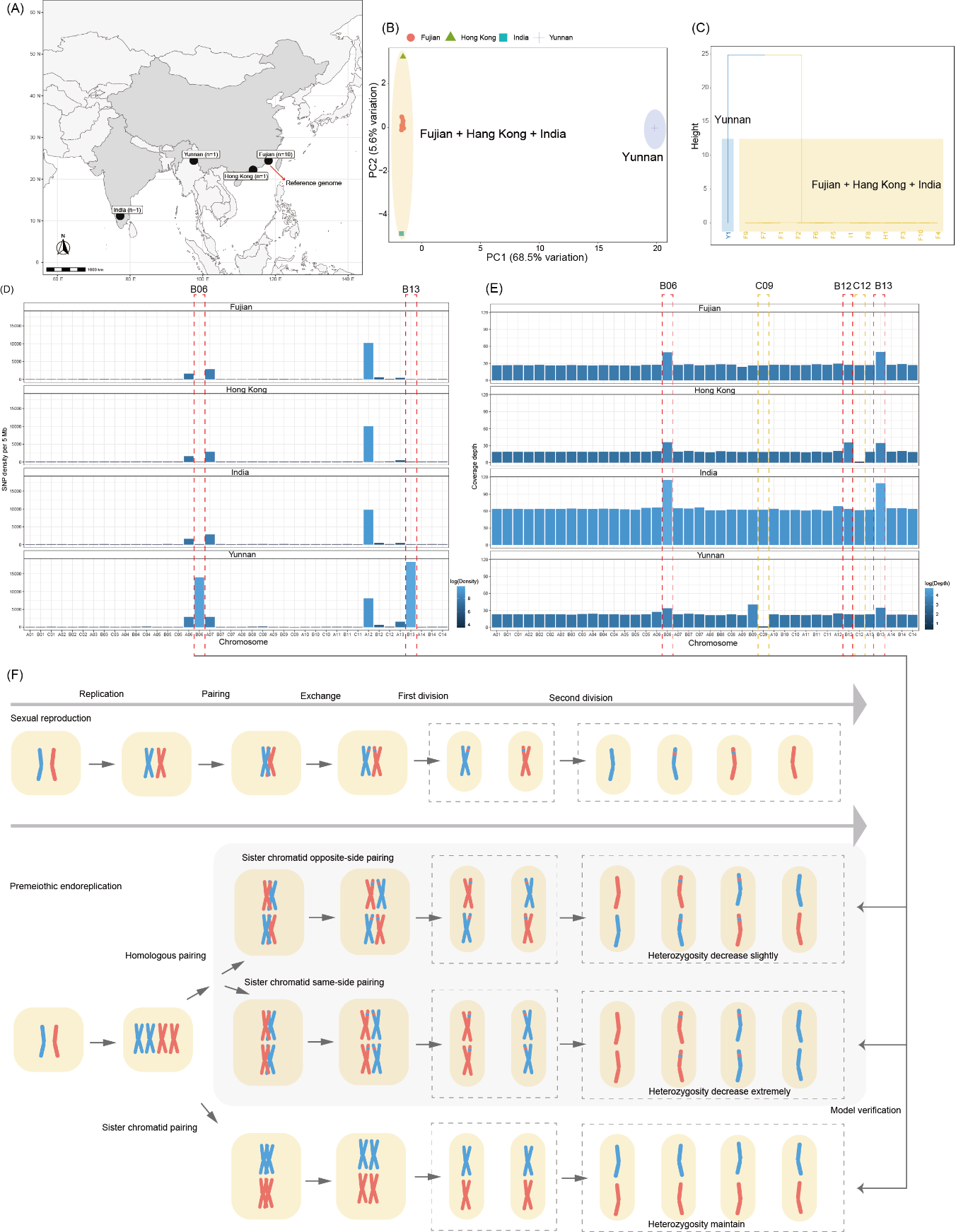
Figure 3. Proposed mechanism of premeiotic endoreplication facilitating parthenogenetic reproduction in the flowerpot snake.
This study not only deepens our understanding of reptilian genome evolution but also challenges long-held assumptions regarding the limitations of asexual reproduction. As researchers continue to explore the genomic underpinnings of parthenogenesis, the flowerpot snake may serve as a key model for studying evolutionary innovation in vertebrates.
Source. https://doi.org/10.1126/sciadv.adt6477